This HTML version of "Think Stats 2e" is provided for convenience, but it is not the best format for the book. In particular, some of the math symbols are not rendered correctly.
You might prefer to read the PDF version, or you can buy a hard copy from Amazon.
Chapter 6 Probability density functions
The code for this chapter is in density.py. For information
about downloading and working with this code, see Section ??.
6.1 PDFs
The derivative of a CDF is called a probability density function, or PDF. For example, the PDF of an exponential distribution is
| PDFexpo(x) = λ e−λ x |
The PDF of a normal distribution is
| PDFnormal(x) = |
| exp | ⎡ ⎢ ⎢ ⎢ ⎢ ⎢ ⎣ | − |
| ⎛ ⎜ ⎜ ⎝ |
| ⎞ ⎟ ⎟ ⎠ |
| ⎤ ⎥ ⎥ ⎥ ⎥ ⎥ ⎦ |
Evaluating a PDF for a particular value of x is usually not useful. The result is not a probability; it is a probability density.
In physics, density is mass per unit of volume; in order to get a mass, you have to multiply by volume or, if the density is not constant, you have to integrate over volume.
Similarly, probability density measures probability per unit of x. In order to get a probability mass, you have to integrate over x.
thinkstats2 provides a class called Pdf that represents
a probability density function. Every Pdf object provides the
following methods:
Density, which takes a value,x, and returns the density of the distribution atx.Render, which evaluates the density at a discrete set of values and returns a pair of sequences: the sorted values,xs, and their probability densities,ds.MakePmf, which evaluatesDensityat a discrete set of values and returns a normalized Pmf that approximates the Pdf.GetLinspace, which returns the default set of points used byRenderandMakePmf.
Pdf is an abstract parent class, which means you should not
instantiate it; that is, you cannot create a Pdf object. Instead, you
should define a child class that inherits from Pdf and provides
definitions of Density and GetLinspace. Pdf provides
Render and MakePmf.
For example, thinkstats2 provides a class named NormalPdf that
evaluates the normal density function.
class NormalPdf(Pdf):
def __init__(self, mu=0, sigma=1, label=''):
self.mu = mu
self.sigma = sigma
self.label = label
def Density(self, xs):
return scipy.stats.norm.pdf(xs, self.mu, self.sigma)
def GetLinspace(self):
low, high = self.mu-3*self.sigma, self.mu+3*self.sigma
return np.linspace(low, high, 101)
The NormalPdf object contains the parameters mu and
sigma. Density uses
scipy.stats.norm, which is an object that represents a normal
distribution and provides cdf and pdf, among other
methods (see Section ??).
The following example creates a NormalPdf with the mean and variance of adult female heights, in cm, from the BRFSS (see Section ??). Then it computes the density of the distribution at a location one standard deviation from the mean.
>>> mean, var = 163, 52.8
>>> std = math.sqrt(var)
>>> pdf = thinkstats2.NormalPdf(mean, std)
>>> pdf.Density(mean + std)
0.0333001
The result is about 0.03, in units of probability mass per cm. Again, a probability density doesn’t mean much by itself. But if we plot the Pdf, we can see the shape of the distribution:
>>> thinkplot.Pdf(pdf, label='normal')
>>> thinkplot.Show()
thinkplot.Pdf plots the Pdf as a smooth function,
as contrasted with thinkplot.Pmf, which renders a Pmf as a
step function. Figure ?? shows the result, as well
as a PDF estimated from a sample, which we’ll compute in the next
section.
You can use MakePmf to approximate the Pdf:
>>> pmf = pdf.MakePmf()
By default, the resulting Pmf contains 101 points equally spaced from
mu - 3*sigma to mu + 3*sigma. Optionally, MakePmf
and Render can take keyword arguments low, high,
and n.
Figure 6.1: A normal PDF that models adult female height in the U.S., and the kernel density estimate of a sample with n=500.
6.2 Kernel density estimation
Kernel density estimation (KDE) is an algorithm that takes a sample and finds an appropriately smooth PDF that fits the data. You can read details at http://en.wikipedia.org/wiki/Kernel_density_estimation.
scipy provides an implementation of KDE and thinkstats2
provides a class called EstimatedPdf that uses it:
class EstimatedPdf(Pdf):
def __init__(self, sample):
self.kde = scipy.stats.gaussian_kde(sample)
def Density(self, xs):
return self.kde.evaluate(xs)
__init__ takes a sample
and computes a kernel density estimate. The result is a
gaussian_kde object that provides an evaluate
method.
Density takes a value or sequence, calls
gaussian_kde.evaluate, and returns the resulting density. The
word “Gaussian” appears in the name because it uses a filter based
on a Gaussian distribution to smooth the KDE.
Here’s an example that generates a sample from a normal distribution and then makes an EstimatedPdf to fit it:
>>> sample = [random.gauss(mean, std) for i in range(500)]
>>> sample_pdf = thinkstats2.EstimatedPdf(sample)
>>> thinkplot.Pdf(sample_pdf, label='sample KDE')
sample is a list of 500 random heights.
sample_pdf is a Pdf object that contains the estimated
KDE of the sample.
Figure ?? shows the normal density function and a KDE based on a sample of 500 random heights. The estimate is a good match for the original distribution.
Estimating a density function with KDE is useful for several purposes:
- Visualization: During the exploration phase of a project, CDFs are usually the best visualization of a distribution. After you look at a CDF, you can decide whether an estimated PDF is an appropriate model of the distribution. If so, it can be a better choice for presenting the distribution to an audience that is unfamiliar with CDFs.
- Interpolation: An estimated PDF is a way to get from a sample to a model of the population. If you have reason to believe that the population distribution is smooth, you can use KDE to interpolate the density for values that don’t appear in the sample.
- Simulation: Simulations are often based on the distribution of a sample. If the sample size is small, it might be appropriate to smooth the sample distribution using KDE, which allows the simulation to explore more possible outcomes, rather than replicating the observed data.
6.3 The distribution framework
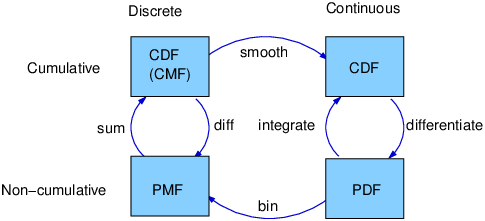
Figure 6.2: A framework that relates representations of distribution functions.
At this point we have seen PMFs, CDFs and PDFs; let’s take a minute to review. Figure ?? shows how these functions relate to each other.
We started with PMFs, which represent the probabilities for a discrete set of values. To get from a PMF to a CDF, you add up the probability masses to get cumulative probabilities. To get from a CDF back to a PMF, you compute differences in cumulative probabilities. We’ll see the implementation of these operations in the next few sections.
A PDF is the derivative of a continuous CDF; or, equivalently, a CDF is the integral of a PDF. Remember that a PDF maps from values to probability densities; to get a probability, you have to integrate.
To get from a discrete to a continuous distribution, you can perform various kinds of smoothing. One form of smoothing is to assume that the data come from an analytic continuous distribution (like exponential or normal) and to estimate the parameters of that distribution. Another option is kernel density estimation.
The opposite of smoothing is discretizing, or quantizing. If you evaluate a PDF at discrete points, you can generate a PMF that is an approximation of the PDF. You can get a better approximation using numerical integration.
To distinguish between continuous and discrete CDFs, it might be better for a discrete CDF to be a “cumulative mass function,” but as far as I can tell no one uses that term.
6.4 Hist implementation
At this point you should know how to use the basic types provided
by thinkstats2: Hist, Pmf, Cdf, and Pdf. The next few sections
provide details about how they are implemented. This material
might help you use these classes more effectively, but it is not
strictly necessary.
Hist and Pmf inherit from a parent class called _DictWrapper.
The leading underscore indicates that this class is “internal;” that
is, it should not be used by code in other modules. The name
indicates what it is: a dictionary wrapper. Its primary attribute is
d, the dictionary that maps from values to their frequencies.
The values can be any hashable type. The frequencies should be integers, but can be any numeric type.
_DictWrapper contains methods appropriate for both
Hist and Pmf, including __init__, Values,
Items and Render. It also provides modifier
methods Set, Incr, Mult, and Remove. These
methods are all implemented with dictionary operations. For example:
# class _DictWrapper
def Incr(self, x, term=1):
self.d[x] = self.d.get(x, 0) + term
def Mult(self, x, factor):
self.d[x] = self.d.get(x, 0) * factor
def Remove(self, x):
del self.d[x]
Hist also provides Freq, which looks up the frequency
of a given value.
Because Hist operators and methods are based on dictionaries, these methods are constant time operations; that is, their run time does not increase as the Hist gets bigger.
6.5 Pmf implementation
Pmf and Hist are almost the same thing, except that a Pmf maps values to floating-point probabilities, rather than integer frequencies. If the sum of the probabilities is 1, the Pmf is normalized.
Pmf provides Normalize, which computes the sum of the
probabilities and divides through by a factor:
# class Pmf
def Normalize(self, fraction=1.0):
total = self.Total()
if total == 0.0:
raise ValueError('Total probability is zero.')
factor = float(fraction) / total
for x in self.d:
self.d[x] *= factor
return total
fraction determines the sum of the probabilities after
normalizing; the default value is 1. If the total probability is 0,
the Pmf cannot be normalized, so Normalize raises ValueError.
Hist and Pmf have the same constructor. It can take
as an argument a dict, Hist, Pmf or Cdf, a pandas
Series, a list of (value, frequency) pairs, or a sequence of values.
If you instantiate a Pmf, the result is normalized. If you instantiate a Hist, it is not. To construct an unnormalized Pmf, you can create an empty Pmf and modify it. The Pmf modifiers do not renormalize the Pmf.
6.6 Cdf implementation
A CDF maps from values to cumulative probabilities, so I could have
implemented Cdf as a _DictWrapper. But the values in a CDF are
ordered and the values in a _DictWrapper are not. Also, it is
often useful to compute the inverse CDF; that is, the map from
cumulative probability to value. So the implementaion I chose is two
sorted lists. That way I can use binary search to do a forward or
inverse lookup in logarithmic time.
The Cdf constructor can take as a parameter a sequence of values or a pandas Series, a dictionary that maps from values to probabilities, a sequence of (value, probability) pairs, a Hist, Pmf, or Cdf. Or if it is given two parameters, it treats them as a sorted sequence of values and the sequence of corresponding cumulative probabilities.
Given a sequence, pandas Series, or dictionary, the constructor makes a Hist. Then it uses the Hist to initialize the attributes:
self.xs, freqs = zip(*sorted(dw.Items()))
self.ps = np.cumsum(freqs, dtype=np.float)
self.ps /= self.ps[-1]
xs is the sorted list of values; freqs is the list
of corresponding frequencies. np.cumsum computes
the cumulative sum of the frequencies. Dividing through by the
total frequency yields cumulative probabilities.
For n values, the time to construct the
Cdf is proportional to n logn.
Here is the implementation of Prob, which takes a value
and returns its cumulative probability:
# class Cdf
def Prob(self, x):
if x < self.xs[0]:
return 0.0
index = bisect.bisect(self.xs, x)
p = self.ps[index - 1]
return p
The bisect module provides an implementation of binary search.
And here is the implementation of Value, which takes a
cumulative probability and returns the corresponding value:
# class Cdf
def Value(self, p):
if p < 0 or p > 1:
raise ValueError('p must be in range [0, 1]')
index = bisect.bisect_left(self.ps, p)
return self.xs[index]
Given a Cdf, we can compute the Pmf by computing differences between
consecutive cumulative probabilities. If you call the Cdf constructor
and pass a Pmf, it computes differences by calling Cdf.Items:
# class Cdf
def Items(self):
a = self.ps
b = np.roll(a, 1)
b[0] = 0
return zip(self.xs, a-b)
np.roll shifts the elements of a to the right, and “rolls”
the last one back to the beginning. We replace the first element of
b with 0 and then compute the difference a-b. The result
is a NumPy array of probabilities.
Cdf provides Shift and Scale, which modify the
values in the Cdf, but the probabilities should be treated as
immutable.
6.7 Moments
Any time you take a sample and reduce it to a single number, that number is a statistic. The statistics we have seen so far include mean, variance, median, and interquartile range.
A raw moment is a kind of statistic. If you have a sample of values, xi, the kth raw moment is:
| m′k = |
|
| xik |
Or if you prefer Python notation:
def RawMoment(xs, k):
return sum(x**k for x in xs) / len(xs)
When k=1 the result is the sample mean, x. The other raw moments don’t mean much by themselves, but they are used in some computations.
The central moments are more useful. The kth central moment is:
| mk = |
|
| (xi − x)k |
Or in Python:
def CentralMoment(xs, k):
mean = RawMoment(xs, 1)
return sum((x - mean)**k for x in xs) / len(xs)
When k=2 the result is the second central moment, which you might recognize as variance. The definition of variance gives a hint about why these statistics are called moments. If we attach a weight along a ruler at each location, xi, and then spin the ruler around the mean, the moment of inertia of the spinning weights is the variance of the values. If you are not familiar with moment of inertia, see http://en.wikipedia.org/wiki/Moment_of_inertia.
When you report moment-based statistics, it is important to think about the units. For example, if the values xi are in cm, the first raw moment is also in cm. But the second moment is in cm2, the third moment is in cm3, and so on.
Because of these units, moments are hard to interpret by themselves. That’s why, for the second moment, it is common to report standard deviation, which is the square root of variance, so it is in the same units as xi.
6.8 Skewness
Skewness is a property that describes the shape of a distribution. If the distribution is symmetric around its central tendency, it is unskewed. If the values extend farther to the right, it is “right skewed” and if the values extend left, it is “left skewed.”
This use of “skewed” does not have the usual connotation of “biased.” Skewness only describes the shape of the distribution; it says nothing about whether the sampling process might have been biased.
Several statistics are commonly used to quantify the skewness of a distribution. Given a sequence of values, xi, the sample skewness, g1, can be computed like this:
def StandardizedMoment(xs, k):
var = CentralMoment(xs, 2)
std = math.sqrt(var)
return CentralMoment(xs, k) / std**k
def Skewness(xs):
return StandardizedMoment(xs, 3)
g1 is the third standardized moment, which means that it has been normalized so it has no units.
Negative skewness indicates that a distribution skews left; positive skewness indicates that a distribution skews right. The magnitude of g1 indicates the strength of the skewness, but by itself it is not easy to interpret.
In practice, computing sample skewness is usually not a good idea. If there are any outliers, they have a disproportionate effect on g1.
Another way to evaluate the asymmetry of a distribution is to look at the relationship between the mean and median. Extreme values have more effect on the mean than the median, so in a distribution that skews left, the mean is less than the median. In a distribution that skews right, the mean is greater.
Pearson’s median skewness coefficient is a measure of skewness based on the difference between the sample mean and median:
| gp = 3 (x − m) / S |
Where x is the sample mean, m is the median, and S is the standard deviation. Or in Python:
def Median(xs):
cdf = thinkstats2.Cdf(xs)
return cdf.Value(0.5)
def PearsonMedianSkewness(xs):
median = Median(xs)
mean = RawMoment(xs, 1)
var = CentralMoment(xs, 2)
std = math.sqrt(var)
gp = 3 * (mean - median) / std
return gp
This statistic is robust, which means that it is less vulnerable to the effect of outliers.
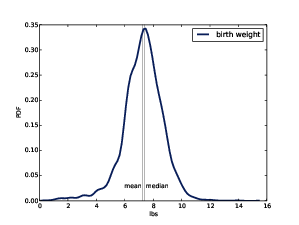
Figure 6.3: Estimated PDF of birthweight data from the NSFG.
As an example, let’s look at the skewness of birth weights in the NSFG pregnancy data. Here’s the code to estimate and plot the PDF:
live, firsts, others = first.MakeFrames()
data = live.totalwgt_lb.dropna()
pdf = thinkstats2.EstimatedPdf(data)
thinkplot.Pdf(pdf, label='birth weight')
Figure ?? shows the result. The left tail appears longer than the right, so we suspect the distribution is skewed left. The mean, 7.27 lbs, is a bit less than the median, 7.38 lbs, so that is consistent with left skew. And both skewness coefficients are negative: sample skewness is -0.59; Pearson’s median skewness is -0.23.
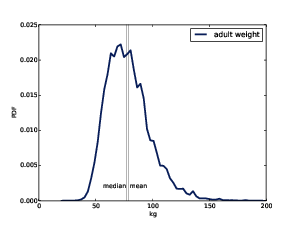
Figure 6.4: Estimated PDF of adult weight data from the BRFSS.
Now let’s compare this distribution to the distribution of adult weight in the BRFSS. Again, here’s the code:
df = brfss.ReadBrfss(nrows=None)
data = df.wtkg2.dropna()
pdf = thinkstats2.EstimatedPdf(data)
thinkplot.Pdf(pdf, label='adult weight')
Figure ?? shows the result. The distribution appears skewed to the right. Sure enough, the mean, 79.0, is bigger than the median, 77.3. The sample skewness is 1.1 and Pearson’s median skewness is 0.26.
The sign of the skewness coefficient indicates whether the distribution skews left or right, but other than that, they are hard to interpret. Sample skewness is less robust; that is, it is more susceptible to outliers. As a result it is less reliable when applied to skewed distributions, exactly when it would be most relevant.
Pearson’s median skewness is based on a computed mean and variance, so it is also susceptible to outliers, but since it does not depend on a third moment, it is somewhat more robust.
6.9 Exercises
A solution to this exercise is in chap06soln.py.
The distribution of income is famously skewed to the right. In this exercise, we’ll measure how strong that skew is.
The Current Population Survey (CPS) is a joint effort of the Bureau
of Labor Statistics and the Census Bureau to study income and related
variables. Data collected in 2013 is available from
http://www.census.gov/hhes/www/cpstables/032013/hhinc/toc.htm.
I downloaded hinc06.xls, which is an Excel spreadsheet with
information about household income, and converted it to hinc06.csv,
a CSV file you will find in the repository for this book. You
will also find hinc2.py, which reads this file and transforms
the data.
The dataset is in the form of a series of income ranges and the number of respondents who fell in each range. The lowest range includes respondents who reported annual household income “Under $5000.” The highest range includes respondents who made “$250,000 or more.”
To estimate mean and other statistics from these data, we have to
make some assumptions about the lower and upper bounds, and how
the values are distributed in each range. hinc2.py provides
InterpolateSample, which shows one way to model
this data. It takes a DataFrame with a column, income, that
contains the upper bound of each range, and freq, which contains
the number of respondents in each frame.
It also takes log_upper, which is an assumed upper bound
on the highest range, expressed in log10 dollars.
The default value, log_upper=6.0 represents the assumption
that the largest income among the respondents is
106, or one million dollars.
InterpolateSample generates a pseudo-sample; that is, a sample
of household incomes that yields the same number of respondents
in each range as the actual data. It assumes that incomes in
each range are equally spaced on a log10 scale.
Compute the median, mean, skewness and Pearson’s skewness of the resulting sample. What fraction of households reports a taxable income below the mean? How do the results depend on the assumed upper bound?
6.10 Glossary
- Probability density function (PDF): The derivative of a continuous CDF, a function that maps a value to its probability density.
- Probability density: A quantity that can be integrated over a range of values to yield a probability. If the values are in units of cm, for example, probability density is in units of probability per cm.
- Kernel density estimation (KDE): An algorithm that estimates a PDF based on a sample.
- discretize: To approximate a continuous function or distribution with a discrete function. The opposite of smoothing.
- raw moment: A statistic based on the sum of data raised to a power.
- central moment: A statistic based on deviation from the mean, raised to a power.
- standardized moment: A ratio of moments that has no units.
- skewness: A measure of how asymmetric a distribution is.
- sample skewness: A moment-based statistic intended to quantify the skewness of a distribution.
- Pearson’s median skewness coefficient: A statistic intended to quantify the skewness of a distribution based on the median, mean, and standard deviation.
- robust: A statistic is robust if it is relatively immune to the effect of outliers.